Submitted on: 2025, 27th January; revised on: 2025, 9th March; accepted on: 2025, 5th May
Genetically informed range extension for Kurixalus inexpectatus (Anura: Rhacophoridae) in Fujian, with ecological niche modeling to guide further searches
1 Laboratory of Animal Behaviour and Conservation, College of Ecology and the Environment, Nanjing Forestry University, Nanjing 210037, China
2 Laboratory of Animal Behaviour and Conservation, College of Life Sciences, Nanjing Forestry University, Nanjing 210037, China
3 College of Life Sciences, Nanjing Forestry University, Nanjing 210037, China
4 Yunxiao County Forestry Bureau, Zhangzhou 363300, China
5 Fujian Forestry Prospect and Design Institute, Fuzhou 350001, China
6 Department of Life Sciences, National Chung Hsing University, Taichung, 402202, Taiwan
7 Global Change Biology Research Center, National Chung Hsing University, Taichung, 402202, Taiwan
8 Department of Zoology and General Biology, Faculty of Life Sciences, Fergana State University, Fergana, 150100, Uzbekistan
*Corresponding authors. Email: dy.othman@gmail.com, amaelborzee@gmail.com
Abstract. Undocumented and unnamed biodiversity is difficult to conserve effectively. We encountered several Kurixalus (Anura: Rhacophoridae) individuals of an unknown species identity in Fujian, China, a province with no previous records of Kurixalus. This genus of frogs, of which many new species have been described in recent years, has been the subject of much historical taxonomic confusion, largely due to conserved morphology. We sequenced two mitochondrial and one nuclear genes (1748 total bp) for three individuals from Fujian and reconstructed their phylogeny including all known lineages of Kurixalus from mainland China, and other East Asia lineages for which data were available. Both Bayesian Inference and Maximum Likelihood trees consistently showed that the individuals from Fujian were most closely related to K. inexpectatus, which previously was only known from northern Zhejiang, over 830 km north of this new location. Our results help clarify the identity of the unknown Kurixalus in Fujian and the distribution of Kurixalus, particularly regarding the distribution of K. hainanus, which has previously been confused with other members of the K. odontotarsus species complex. Additionally, we created ecological niche models using Maxent for the clade including both K. inexpectatus and its sister species K. idiootocus to guide future survey efforts.
Keywords. Kurixalus, Maxent, cryptic species, tree frog.
The genus Kurixalus Ye, Fei, and Dubois, 1999, sometimes referred to as the frilled swamp tree frogs, consists of 23 currently recognized species, with ten of these described since 2014 (Frost, 2024a). These small, arboreal frogs of the family Rhacophoridae are found mainly in Southeast Asia and have relatively similar morphologies, which has led to significant taxonomic confusion within the genus (Yu et al., 2017a; Lv et al., 2018; Nguyen et al., 2020). Kurixalus has also been the subject of multiple studies of biogeography and patterns of historical dispersal between islands and the Asian mainland (Yu et al., 2020; Mo et al., 2023), although some reported colonization timelines dating to the Miocene are incompatible with the estimated timing of geological formation of landmasses in the Pliocene (Lv et al., 2018; Ali, 2020).
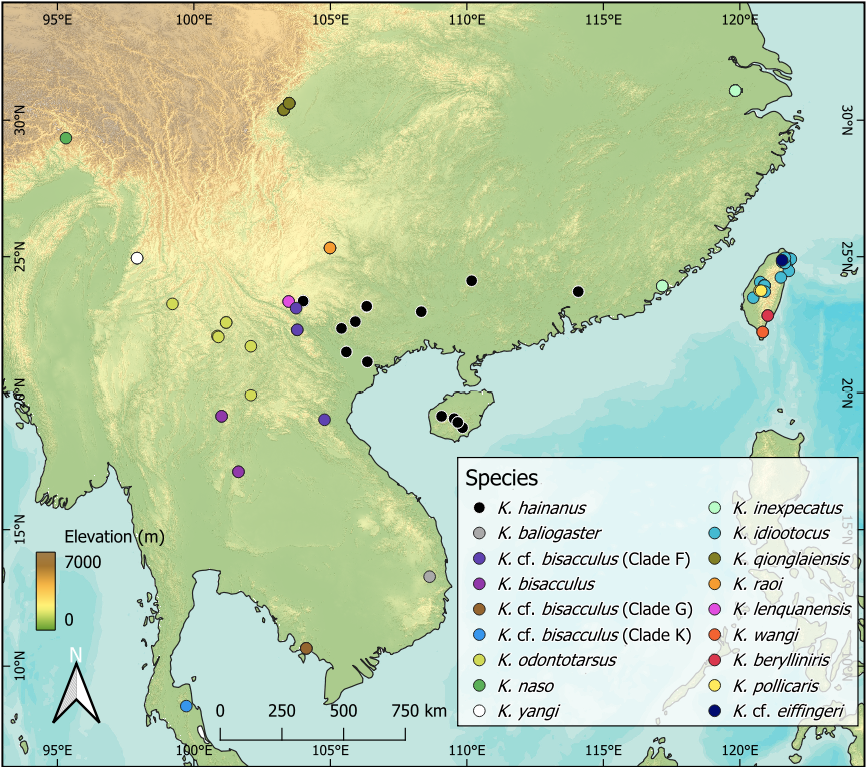
Several recently described species of Kurixalus are known from only the type locality or very small ranges (Yu et al., 2018; Hou et al., 2021; Zeng et al., 2021; Guo et al., 2022; Messenger et al., 2022), but additional surveys may reveal new localities, as was the case with K. lenquanensis (Yu et al., 2017b; Pang et al., 2024). During field surveys at night in February 2024 in Yunxiao County (Fujian, China), we encountered multiple individuals of the genus Kurixalus of uncertain species identity (Fig. 1). The frogs were calling from underneath vegetation in a montane wetland at an elevation of around 800 meters (23.9189° N, 117.2022° E). As far as we are aware, there have been no published records for the genus Kurixalus in Fujian, with the closest records on the Chinese mainland being in Guangdong and belonging to K. hainanus (Yu et al., 2017a). Given the presence of paired dark patches on the belly, we hypothesized the unknown Kurixalus were most likely related to K. idiootocus or K. inexpectatus rather than K. hainanus (Zeng et al., 2021; Messenger et al., 2022). The K. idiootocus species complex has been in flux recently, with five mainland species having been described as sister to the island endemic K. idiootocus, namely K. lenquanensis (Yu et al., 2017b), K. raoi (Zeng et al., 2021), K. silvaenaias (Hou et al., 2021), K. qionglaiensis (Guo et al., 2022), and K. inexpectatus (Messenger et al., 2022). Of these, K. silvaenaias and K. qionglaiensis have been recognized to be the same species and the synonymy of K. inexpectatus with K. idiootocus has been proposed as well (Lyu et al., 2024). Additionally, the distribution and identity of members of the K. odontotarsus species complex in Southeast Asia is also in need of further clarification. Several likely species-level lineages have been identified (Yu et al., 2017a) but only some of these have been formally described, such as K. yangi (Yu et al., 2018). Within southeastern China, specifically Guangxi, Guangdong, and Hainan, the names K. bisacculus, K. hainanus, K. odontotarsus, and K. verrucosus have been used, but they likely correspond to a single lineage within the K. odontotarsus species complex (Yu et al., 2017a; Lv et al., 2018; Mo et al., 2023).

We conducted molecular phylogenetic analyses to assess the identity of the unidentified frogs and clarify the distribution of Kurixalus frogs in eastern Asia, with an emphasis on mainland China. Preliminary molecular barcoding indicated that the specimens from Fujian were most closely related to K. inexpectatus, though the only previously known locality of K. inexpectatus is in northern Zhejiang, 830 km further north. We also used ecological niche modeling with the known points for K. inexpectatus and its sister species, K. idiootocus, to predict the areas that may harbor additional undiscovered populations.
Upon encounter, three Kurixalus individuals from Fujian were collected and buccal swabs were taken to obtain DNA. We extracted genomic DNA from the buccal swabs of the three Kurixalus individuals sampled in Fujian and three individuals of K. idiootocus from Taiwan, as well as DNA from thigh muscle of four specimens of K. inexpectatus from Zhejiang (Table S1) using a Qiagen DNeasy Blood & Tissue Kit (Qiagen, Germany) according to the manufacturer’s protocol. For the Kurixalus individuals from Fujian and three K. idiootocus from Taiwan, we sequenced one nuclear and two mitochondrial fragments. For the nuclear gene, we sequenced a 476 bp long fragment of Tyrosine exon-1 (TYR) using the primer pair L2976 (5’-TGC TGG GCR TCT CTC CAR TCC CA-3’) H2977 (5’-AGG TCC TCY TRA GGA AGG AAT G-3’; Bossuyt and Milinkovitch, 2000). For the mtDNA, we sequenced an 827 bp fragment from a section of the partial 12S rRNA, complete tRNA-Valine, and partial 16S rRNA (12S-Val-16S) genes using the primer pair F0001 (5’-AGA TAC CCC ACT ATG CCT ACC C-3’) R1169 (5’-GTG GCT GCT TTT AGG CCC ACT-3’; Wilkinson, Drewes, and Tatum, 2002). We also sequenced a 554 bp long fragment of the cytochrome oxidase subunit I (COI) gene using the primer pairs COI-CO1 (5’-TYT CWA CWA AYC AYA AAG AYA TTG G-3’) COI-CO3 (5’-ACY TCY GGR TGA CCA AAR AAY CA-3’) and Chmf4 (5’-TYT CWA CWA AYC AYA AAG AYA TCG G-3’) Chmr4 (5’-ACY TCR GGR TGR CCR AAR AAT CA-3’; Che et al., 2012). For the K. inexpectatus from Zhejiang, which already had two gene fragments sequenced (Messenger et al., 2022), we sequenced COI using the ad hoc-designed primers CO1KuF (5’-CCT GGG CCG GAA TGA TCG-3’) CO1KuR (5’-TTG ATA AAG AAC TGG GTC CCC-3’), as these samples failed to amplify with the COI primers mentioned above. We amplified all fragments using polymerase chain reactions (PCR) in a total volume of 25 μl, which contained 12.5 μl of 2× Hieff PCR Master Mix (without dye), 1 μl of a 10 μM solution of each primer, 2 μl of DNA sample at a concentration of 10 ng/μl (within the recommended range), and 8.5 μl ddH2O. We carried out amplification using an Arhat 96 thermal cycler (Shanghai, China). Thermal profiles for PCR were as follows: initial denaturation at 95 °C for 5 minutes, followed by 35 cycles of denaturation at 94 °C for 1 minute, annealing at 54 °C for TYR, 55 °C for 12S-Val-16S, and 46 °C for COI for 1 minute, and extension at 72 °C for 1 minute. The cycles were followed by a 10-minute final extension at 72 °C. PCR amplifications and double reads sequencing for all samples were performed by Sangon Biotech Co., Ltd. (Shanghai, China) and Tsingke Biotech Co., Ltd. (Beijing, China). The Kurixalus individuals from Fujian were released after initial DNA barcoding indicated they were not an undescribed species and no morphological measurements were taken.
To complement our sequences and reconstruct alignments for phylogeny, we downloaded homologous sequences for Kurixalus species from GenBank (www.ncbi.nlm.nih.gov/genbank). We added a sequence of Theloderma albopunctatum (Rhacophoridae) as the outgroup. To identify our unknown samples using phylogenetic analyses, the sequences in our alignments covered all identified clades of Kurixalus found in mainland China (Fig. 2, Table S1), according to the literature (Wilkinson, Drewes, and Tatum, 2002; Frost et al., 2006; Li et al., 2008, 2013; Nguyen, Matsui, and Duc, 2014; Wu et al., 2016; Yu et al., 2017a, 2020; Hou et al., 2021; Zeng et al., 2021; Messenger et al., 2022; Luo et al., 2023; Lyu et al., 2024; Xu et al., 2024). We trimmed and aligned our sequences using Muscle v. 5.1 (Edgar, 2004) in Geneious Prime 2023.2.1 (Kearse et al. 2012; www.geneious.com). We constructed three different sequence alignment datasets, namely: (1) a 1748 bp long concatenation of three gene fragments (802 bp of partial 12S-Val-16S, 553 bp of COI, and 393 bp of TYR), including four individual K. inexpectatus, three K. idiootocus, and the three Kurixalus sampled in Fujian as the ingroup taxa, and two outgroups (K. cf. bisacculus and T. albopunctatum); (2) a 553 bp long COI fragment composed of 91 Kurixalus individuals and one T. albopunctatum, which had the greatest taxonomic coverage of our datasets; and (3) an 802 bp long fragment of 12S-Val-16S composed of 43 Kurixalus individuals and one T. albopunctatum, which had a balance of taxonomic coverage and fragment length.
We used Partition Finder v. 2.1.1 (Guindon et al., 2010; Lanfear et al., 2012, 2016) to determine the best partitioning of the defined subsets, considering a fixed model for non-coding fragments and one for every codon position with respect to the coding fragments. We selected the models based on corrected Akaike information criterion (AICc) values. All models were treated as priors and used for further phylogenetic analyses, which were conducted using Bayesian inference (BI) implemented in MrBayes v. 3.2.4 (Ronquist et al., 2012) and maximum-likelihood (ML) implemented in IQ-Tree (iqtree.cibiv.univie.ac.at; Chernomor et al. 2016; Trifinopoulos et al. 2016; Hoang et al. 2018). For the BI analysis for each dataset, four independent runs were conducted, each of which was performed with MCMC algorithm for 10 million generations. We sampled every 1,000 generations, and discarded the first 25% of samples as burn-in. To ensure the convergence of all runs, we ensured that the analyses reached split frequencies below 0.005. For the ML analysis, we generated consensus trees using 1,000 bootstrap replicates.
Ecological niche modeling, most commonly implemented using Maxent (Phillips et al., 2017), relies on occurrence and environmental data to predict suitable areas for species (Ananjeva et al., 2015; Hou et al., 2023). Such models can be used to guide field surveys for uncommon species (Rhoden et al., 2017; Sarker et al., 2019; Entiauspe‐Neto et al., 2024) and aid in conservation planning (Kidov and Litvinchuk, 2021; Shin et al., 2021). We used Maxent within R v. 4.2.1 to predict what other areas in eastern Asia may harbor undiscovered populations of Kurixalus related to those we found in Fujian (Phillips et al., 2017; Rhoden et al., 2017; R Core Team, 2022). As barcoding suggested, the individuals in Fujian were most closely related to K. inexpectatus, we used occurrence records of K. inexpectatus (Messenger et al., 2022; Li, 2023). However, since three localities is insufficient for accurate species distribution modeling (van Proosdij et al., 2016), we also opted to include an initial 29,251 records from GBIF for K. idiootocus (GBIF.org, 2024b), its sister species (Messenger et al., 2022). We removed the duplicates and thinned the datapoints at a distance of 1 km using the thinData function of the SDMtune package (Vignali et al., 2020), resulting in 1,585 total occurrence points for model training.
We defined the area for model construction as a 50 km buffer around the occurrence records (Hijmans, 2024). We started with the standard 19 bioclimatic variables as climate data (Fick and Hijmans, 2017) and supplemented them with additional layers for elevation, slope, and tree cover (Farr et al., 2007; Zanaga et al., 2021) via the geodata package v.0.6.2 (Hijmans et al., 2024), all at a resolution of 30 arc seconds (approx. 1 km2). We used the built-in GRASS plug-in in QGIS v.3.32.2 (QGIS.org, 2023) to calculate Pearson correlation coefficients for our environmental variables within the study area and identify highly correlated (r > |0.8|) variables (Elith et al., 2011) to remove, though this step is arguably unnecessary (Feng et al., 2019). The following variables were used for modeling: Bio 1 (annual mean temperature), Bio 2 (mean diurnal range), Bio 3 (isothermality), Bio 5 (maximum temperature of warmest month), Bio 12 (annual precipitation), Bio 13 (precipitation of wettest month) and Bio 17 (precipitation of driest quarter), as well as elevation, slope, and tree cover. A total of 10,000 background points were selected from the study area, with selection weighted using a bias raster file generated from 539,243 records of anurans from GBIF within 115.8° to 125.5° latitude and 19.9° to 35.1° longitude (GBIF.org, 2024a) to reduce spatial bias in the occurrence data set (Kramer-Schadt et al., 2013; Zhu and Qiao, 2016).
Candidate Maxent models were generated using ENMeval (Kass et al., 2021) using 30 combinations of feature classes (“L”, “LQ”, “H”, “LQH”, “LQHP”, “LQHPT”) and regularization multipliers (1-5), with data partitioned using ‘checkerboard2’. We selected the model with the lowest AICc, which balances fit and complexity (Warren and Seifert, 2011), as the best model and projected to southeastern Asia. To evaluate model accuracy, we calculated the Boyce index and area under the receiver operating characteristic curve (AUC; Breiner et al., 2015; Liu et al., 2024).
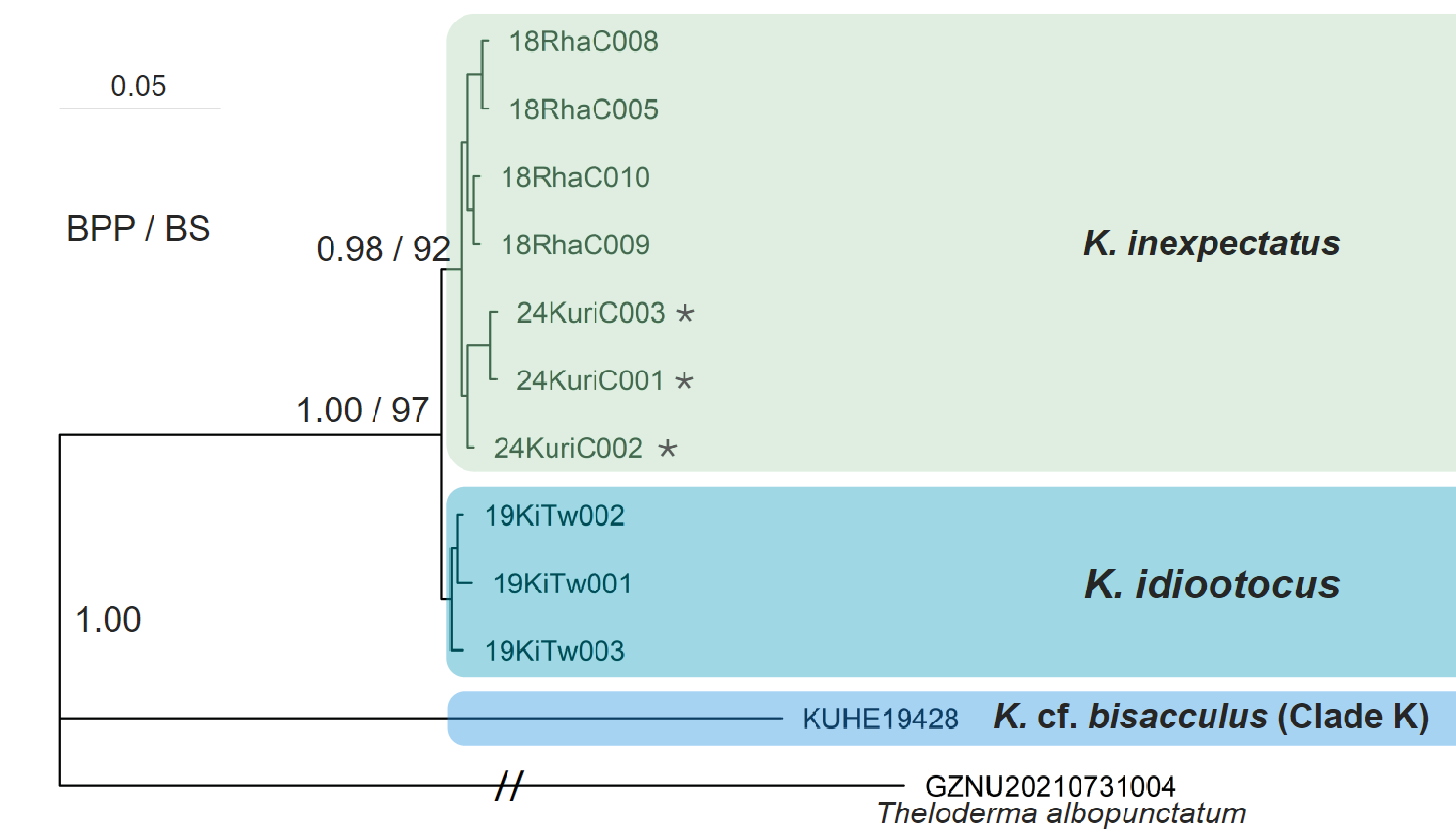
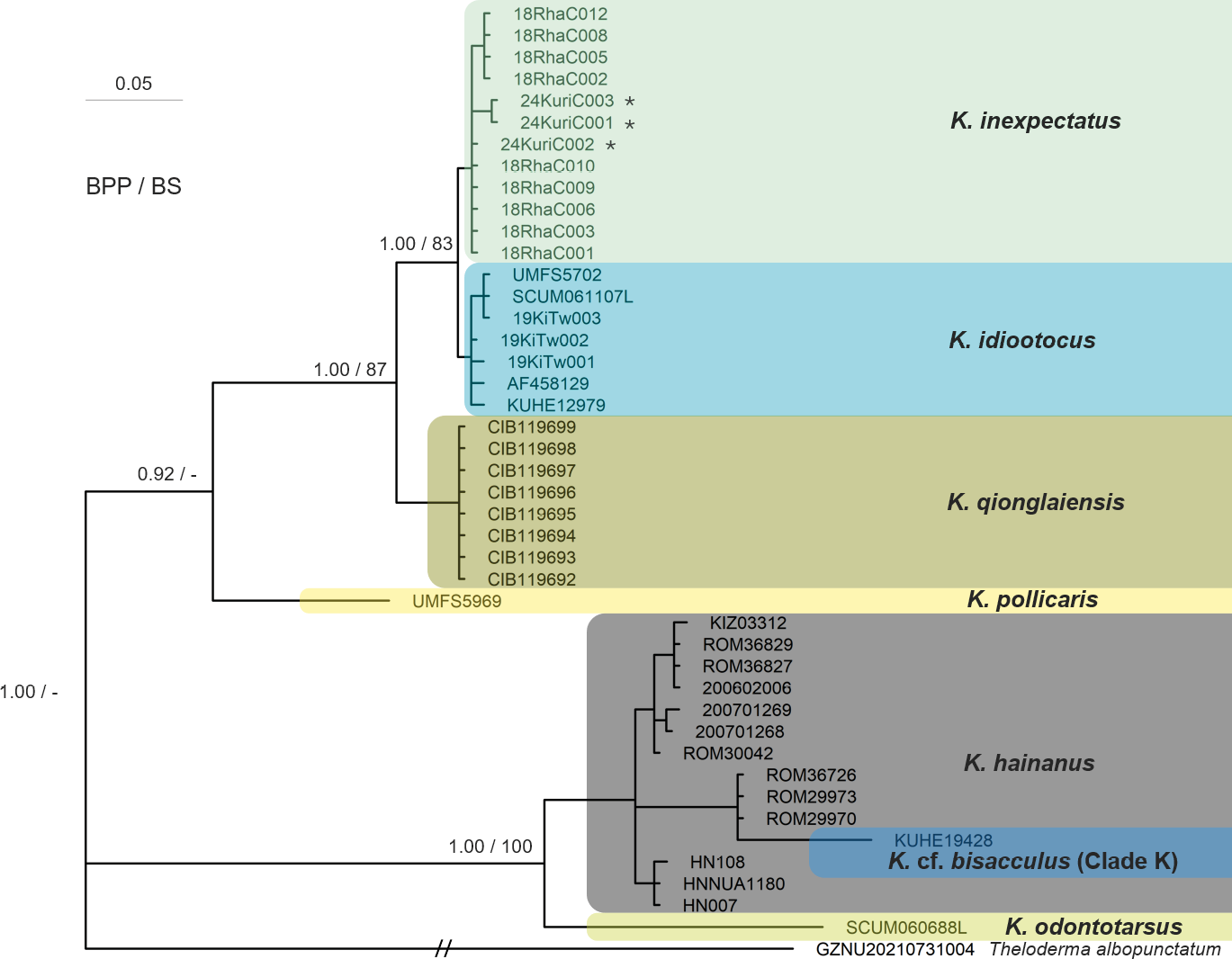
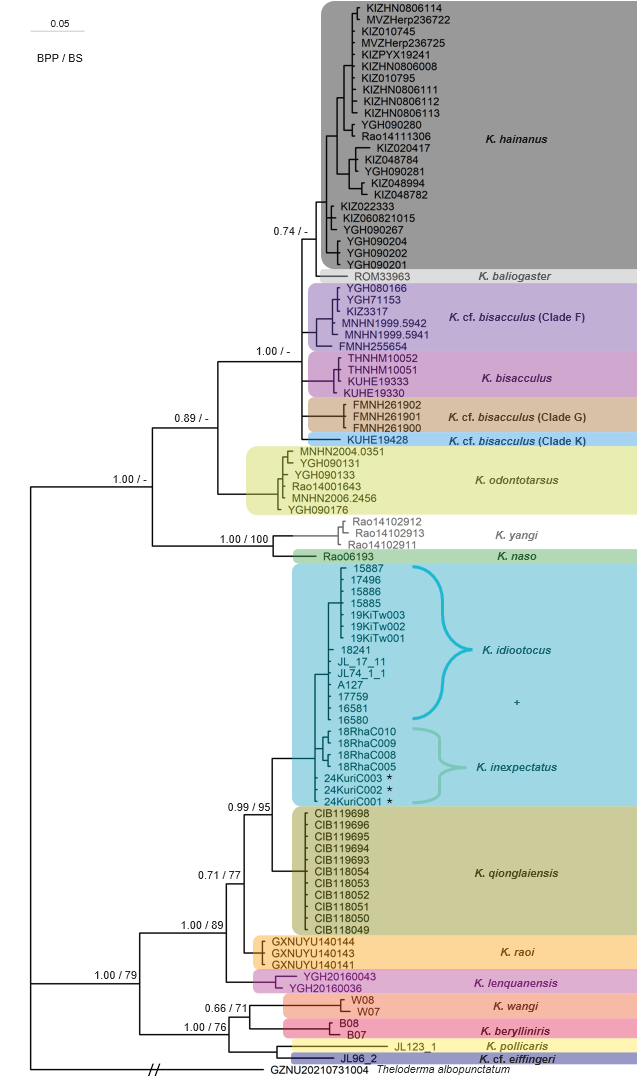
Though the BI and ML trees produced some inconsistencies, in all reconstructed trees the three individuals from Fujian were consistently grouped within a clade of K. idiootocus and K. inexpectatus (Figs. 3-5). Specifically, phylogenetic trees based on the three concatenated gene fragments (Fig. 3) and independent 12S-Val-16S (Fig. 4) datasets indicated the Fujian individuals formed a monophyletic group with K. inexpectatus (Bayesian posterior probability (BPP) = 1.00 and 1.00; ML bootstrap: 97% and 83%, respectively). The tree reconstructed from independent COI fragments (Fig. 5) showed an unclear resolution of this group, forming polytomies for the Fujian samples, K. idiootocus, and K. inexpectatus. According to the BI trees, the group of K. idiootocus, K. inexpectatus, K. raoi, K. lenquanensis, and K. qionglaiensis formed a monophyletic group (BPP = 1.00) which was sister to a clade of island endemics consisting of K. wangi, K. berylliniris, K. pollicaris, and K. cf. eiffingeri (BPP = 1.00). The taxonomy for species names following Frost (2024a) as of July 2024, with unnamed clades following naming from Yu et al. (2017a). In addition, the remaining Kurixalus samples within the K. odontotarsus species group formed a strongly supported clade (BPP = 1.00). Samples from Guangxi, Guangdong, Hainan, and northern Vietnam clustered within the K. odontotarsus species complex but distinct from both K. bisacculus sensu stricto and K. odontotarsus sensu stricto. Other undescribed clades (i.e. K. cf. bisacculus Clades F, G, and K) previously identified in the K. odontotarsus species complex (Yu et al., 2017a) clustered together, but showed unclear resolution between each other.
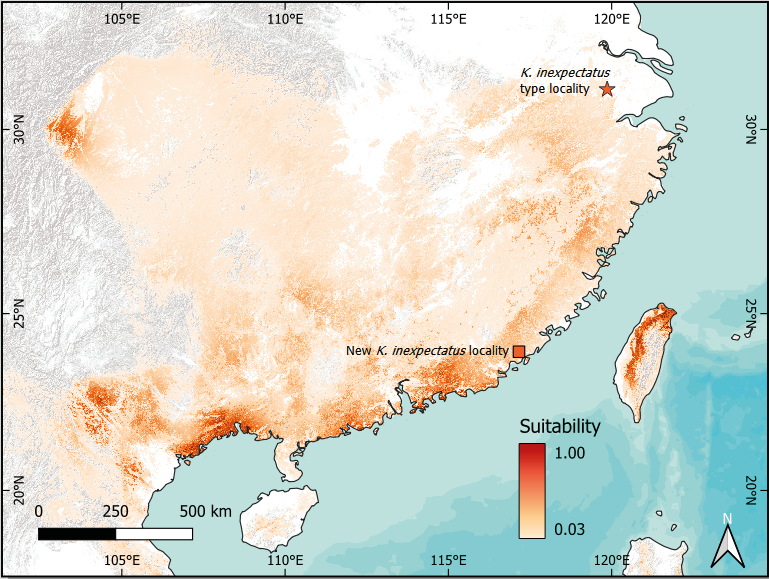
Out of 30 candidate models, the species distribution model for the combined clade of K. inexpectatus and K. idiootocus (Fig. 6) with the lowest AICc used LQHPT feature classes and a regularization multiplier of one. The AUC value was moderately high (0.80), while the Boyce index value was very high (0.98). The highly suitable habitat predicted in Taiwan matched the known distribution of K. idiootocus well, and the model also predicted broad areas of potentially suitable habitat across mainland Southern China and Vietnam. Additional highly suitable areas on the Asian mainland were in northern Vietnam, southern Guangxi, central Sichuan, and along the coast of Guangdong.
Our previously unknown samples from Fujian clustered with K. inexpectatus, which represents a range extension of over 830 km for the species. A previous analysis based on only mtDNA proposed that K. inexpectatus be synonymized with K. idiootocus (Lyu et al., 2024). We acknowledge that the morphological differences between the two species are minor, and that vocalizations within the genus can vary by context and deserve to be studied in further detail (Zhu et al., 2017; Deng et al., 2024; Lyu et al., 2024). Our concatenated tree of 12S-Val-16S, COI, and TYR showed K. idiootocus to be definitively monophyletic, not paraphyletic with respect to K. inexpectatus as previously suggested (Lyu et al., 2024), with high BPP (1.00) for the split between the two. Approaches utilizing more loci and longer fragments are more accurate for species delimitation (Blair and Bryson, 2017; Hofmann et al., 2019; Chan et al., 2022), and previous calls for synonymization with K. idiootocus were based on only one mitochondrial fragment from K. inexpectatus (Lyu et al., 2024). The divergence between K. idiootocus and K. inexpectatus shown in our concatenated tree is shallow though, and the single gene COI tree did not show two reciprocally monophyletic lineages in this clade yet did delineate other known lineages. An integrated taxonomic approach using both morphological measurements and genome-level molecular data would be helpful to clarify the species status of K. inexpectatus. At present, we consider K. inexpectatus to be a valid species, currently known from only northern Zhejiang and southern Fujian (Fig. 2).
Geographically, the closest mainland population to the K. inexpectatus found in Fujian is K. hainanus (Fig. 2), though these two congeners are not closely related. Populations of K. hainanus in Guangdong, Guangxi, and Hainan, and northeastern Vietnam have been previously referred to as K. bisacculus and K. odontotarsus (Yu et al., 2017a; Lv et al., 2018; Mo et al., 2023), but based on the COI tree these individuals form a monophyletic clade, distinct from both K. bisacculus sensu stricto, which is not found in China, and K. odontotarsus sensu stricto, whose presence in China is limited to Yunnan (Yu et al., 2017a). Additional undescribed lineages of the K. odontotarsus species complex exist in Yunnan and Southeast Asia (Yu et al., 2017a, 2020; Frost, 2024b), and significant further work remains to be done in the genus, such as regarding the status of K. pollicaris and K. cf. eifffingeri (Dufresnes and Litvinchuk, 2022). Though it is unknown if K. inexpectatus and K. hainanus overlap in distribution, they can be easily distinguished by the presence (K. inexpectatus and K. idiootocus) or absence (K. hainanus) of paired symmetric dark blotches on the chest (Zhao et al., 2005; Zeng et al., 2021; Messenger et al., 2022).
The results of our modeling indicated broad areas of potentially suitable habitat throughout mainland Southern China for the combined clade of K. inexpectatus and K. idiootocus (Fig. 6). However, it is highly unlikely that K. inexpectatus occurs continuously from northern Zhejiang to southern Fujian, especially given the small range of K. inexpectatus and the other mainland species in the clade (K. qionglaiensis, K. raoi, and K. lenquanensis) and the impact of continued habitat degradation (Pan et al., 2019; Li et al., 2024; Pang et al., 2024). Nevertheless, the existence of additional undocumented populations is possible, particularly in the regions immediately surrounding the two known localities of K. inexpectatus, though surveys near the type locality of the species did not detect it (Kohler et al., 2024). The species has not been evaluated by the IUCN Red List of Threatened Species and estimates of population trends in both known populations of K. inexpectatus are lacking, however, the suitable habitat is decreasing at both localities because of bamboo plantation and the development of infrastructures for tourism (Messenger et al., 2022). Therefore, given the very low extent of occurrence of the species (c. 175 km2) we recommend the species to be listed as endangered under the criteria B1 as the species is present at less than five location (B1a) and there is an observed continuing decline in the quality of the habitat (B1b(iii); IUCN Standards and Petitions Committee, 2024). While habitat loss is the main threat to the species, climate change, pollution are likely to also threaten the species (Luedtke et al., 2023). Interestingly, the highly suitable area forecasted by our model in central Sichuan is actually occupied by the next most closely related species, K. qionglaiensis (Hou et al., 2021; Guo et al., 2022). Similarly, the highly suitable area in southern Guangxi and northern Vietnam is occupied by another congener, K. hainanus. Based on our modeling results and the currently known range of K. inexpectatus, we anticipate two of the more promising areas to search for undiscovered populations to be: (1) near the northern coast of Guangdong; and (2) the northern Wuyi Mountains, Fujian, which is one of the recently identified hotspots of amphibian biodiversity in China (Xu et al., 2024). In addition to our model of broad habitat suitability, future search efforts should also take into account microhabitat and possible breeding conditions conducive to Rhacophorids (Lin and Kam, 2008; Madhushanka and Manamendra-Arachchi, 2021). We are optimistic that additional populations of Kurixalus in mainland China can be discovered with additional searching, as has been the case for other Maxent-guided field efforts (Rhoden, Peterman, and Taylor, 2017; Sarker et al., 2019).
This project was funded by the Research Fund for International Scientists (RFIS) from the National Natural Science Foundation of China (NSFC; W2432021) and the Foreign Youth Talent Program of the Ministry of Science and Technology of the People’s Republic of China (QN2023014004L) awarded to AB. No permits were required for sampling on the mainland, and Kurixalus idiootocus samples were collected under the permit 103-16. This research received the approval from the IACUC Ethics Committee of Nanjing Forestry University, number 2024013.
Supplementary material associated with this article can be found at <http://www9.unipv.it/webshi/appendix> Manuscript number 17227.
Ali, J.R. (2020): Geological data indicate that the interpretation for the age-calibrated phylogeny for the Kurixalus-genus frogs of South, South-east and East Asia (Lv et al., 2018) needs to be rethought. Mol. Phylogenet. Evol. 145: 106053.
Ananjeva, N.B., Golynsky, E.E., Lin, S.-M., Orlov, N.L., Tseng, H.-Y. (2015): Modeling habitat suitability to predict the potential distribution of the Kelung cat snake Boiga kraepelini Steineger, 1902. Russ. J. Herpetol. 22: 197-205.
Blair, C., Bryson, R.W. (2017): Cryptic diversity and discordance in single‐locus species delimitation methods within horned lizards (Phrynosomatidae: Phrynosoma). Mol. Ecol. Resour. 17: 1168-1182.
Bossuyt, F., Milinkovitch, M.C. (2000): Convergent adaptive radiations in Madagascan and Asian ranid frogs reveal covariation between larval and adult traits. Proc. Natl. Acad. Sci. 97: 6585-6590.
Breiner, F.T., Guisan, A., Bergamini, A., Nobis, M.P. (2015): Overcoming limitations of modelling rare species by using ensembles of small models. Methods Ecol. Evol. 6: 1210-1218.
Chan, K.O., Hertwig, S.T., Neokleous, D.N., Flury, J.M., Brown, R.M. (2022): Widely used, short 16S rRNA mitochondrial gene fragments yield poor and erratic results in phylogenetic estimation and species delimitation of amphibians. BMC Ecol. Evol. 22: 37.
Che, J., Chen, H., Yang, J., Jin, J., Jiang, K., Yuan, Z., Murphy, R.W., Zhang, Y. (2012): Universal COI primers for DNA barcoding amphibians. Mol. Ecol. Resour. 12: 247-258.
Chernomor, O., Von Haeseler, A., Minh, B.Q. (2016): Terrace aware data structure for phylogenomic inference from supermatrices. Syst. Biol. 65: 997-1008.
Deng, K., He, Y.-X., Wang, X.-P., Wang, T.-L., Wang, J.-C., Chen, Y.-H., Cui, J.-G. (2024): Hainan frilled treefrogs’ calls partially conform to Menzerath–Altmann’s law, but oppose Zipf’s law of abbreviation. Anim. Behav. 213: 51-59.
Dufresnes, C., Litvinchuk, S.N. (2022): Diversity, distribution and molecular species delimitation in frogs and toads from the Eastern Palaearctic. Zool. J. Linn. Soc. 195: 695-760.
Edgar, R.C. (2004): MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32: 1792-1797.
Elith, J., Phillips, S.J., Hastie, T., Dudík, M., Chee, Y.E., Yates, C.J. (2011): A statistical explanation of MaxEnt for ecologists: Statistical explanation of MaxEnt. Divers. Distrib. 17: 43-57.
Entiauspe‐Neto, O.M., Dervanoski, D., Abegg, A.D. (2024): Can fieldwork driven by predictive species distribution models yield new rare or relevant geographic records? A case study with Neotropical snakes. Austral Ecol. 49: e70013.
Farr, T.G., Rosen, P.A., Caro, E., Crippen, R., Duren, R., Hensley, S., Kobrick, M., Paller, M., Rodriguez, E., Roth, L., Seal, D., Shaffer, S., Shimada, J., Umland, J., Werner, M., Oskin, M., Burbank, D., Alsdorf, D. (2007): The shuttle radar topography mission. Rev. Geophys. 45: 2005RG000183.
Feng, X., Park, D.S., Liang, Y., Pandey, R., Papeş, M. (2019): Collinearity in ecological niche modeling: Confusions and challenges. Ecol. Evol. 9: 10365-10376.
Fick, S.E., Hijmans, R.J. (2017): WorldClim 2: new 1‐km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37: 4302-4315.
Frost, D.R. (2024a): Amphibian species of the world 6.2, an online reference. American Museum of Natural History, New York. Available online at: https://amphibiansoftheworld.amnh.org/. [Accessed 13 September 2024].
Frost, D.R. (2024b): Kurixalus odontotarsus. American Museum of Natural History, New York. Available online at: https://amphibiansoftheworld.amnh.org/Amphibia/Anura/Rhacophoridae/Rhacophorinae/Kurixalus/Kurixalus-odontotarsus. [Accessed 19 November 2024].
Frost, D.R., Grant, T., Faivovich, J., Bain, R.H., Haas, A., Haddad, C.F.B., De Sá, R.O., Channing, A., Wilkinson, M., Donnellan, S.C., Raxworthy, C.J., Campbell, J.A., Blotto, B.L., Moler, P., Drewes, R.C., Nussbaum, R.A., Lynch, J.D., Green, D.M., Wheeler, W.C. (2006): The amphibian tree of life. Bull. Am. Mus. Nat. Hist. 297: 1-291.
GBIF.org (2024a): Occurrence Download: Anura. The Global Biodiversity Information Facility. https://doi.org/10.15468/DL.CXHN8Z.
GBIF.org (2024b): Occurrence Download: Kurixalus idiootocus. The Global Biodiversity Information Facility. https://doi.org/10.15468/DL.XPSDR7.
Guindon, S., Dufayard, J.-F., Lefort, V., Anisimova, M., Hordijk, W., Gascuel, O. (2010): New algorithms and methods to estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 59: 307-321.
Guo, C.-P., Zhong, M.-J., Wah Leung, K., Wang, X.-Y., Hu, J.-H. (2022): A new species of the genus Kurixalus (Anura, Rhacophoridae) from Sichuan Province, southwestern China. Zool. Res. 43: 90-94.
Hijmans, R.J. (2024): raster: Geographic data analysis and modeling.
Hijmans, R.J., Barbosa, M., Ghosh, A., Mandel, A. (2024): ‘geodata’: Download geographic data.
Hoang, D.T., Chernomor, O., Von Haeseler, A., Minh, B.Q., Vinh, L.S. (2018): UFBoot2: Improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35: 518-522.
Hofmann, E.P., Nicholson, K.E., Luque-Montes, I.R., Köhler, G., Cerrato-Mendoza, C.A., Medina-Flores, M., Wilson, L.D., Townsend, J.H. (2019): Cryptic diversity, but to what extent? Discordance between single-locus species delimitation methods within mainland anoles (Squamata: Dactyloidae) of Northern Central America. Front. Genet. 10: 11.
Hou, M., Peng, X., Miao, J., Liu, S., Li, P., Orlov, N.L. (2021): Description a new species of genus Kurixalus (Amphibia: Anura: Rhacophoridae) from Chengdu Prefecture, Sichuan Province, China. Anim. Mol. Breed. 11: 1-16.
Hou, J., Xiang, J., Li, D., Liu, X. (2023): Prediction of potential suitable distribution areas of Quasipaa spinosa in China based on MaxEnt optimization model. Biology 12: 366.
IUCN Standards and Petitions Committee (2024): Guidelines for using the IUCN Red List categories and criteria. Gland, Switzerland.
Kass, J.M., Muscarella, R., Galante, P.J., Bohl, C.L., Pinilla‐Buitrago, G.E., Boria, R.A., Soley‐Guardia, M., Anderson, R.P. (2021): ENMeval 2.0: Redesigned for customizable and reproducible modeling of species’ niches and distributions. Methods Ecol. Evol. 12: 1602-1608.
Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A., Markowitz, S., Duran, C., Thierer, T., Ashton, B., Meintjes, P., Drummond, A. (2012): Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28: 1647-1649.
Kidov, A.A., Litvinchuk, S.N. (2021): Distribution and conservation status of the Hyrcanian Wood Frog (Rana pseudodalmatina) in Azerbaijan. Russ. J. Herpetol. 28: 97-107.
Kohler, D.B., Zhang, X., Messenger, K.R., Chin Yu An, K., Ghosh, D., Othman, S.N., Wang, Z., Amin, H., Prasad, V.K., Wu, Z., Borzée, A. (2024): At home in Jiangsu: Environmental niche modeling and new records for five species of amphibian and reptile in Jiangsu, China. Herpetozoa 37: 85-93.
Kramer-Schadt, S., Niedballa, J., Pilgrim, J.D., Schröder, B., Lindenborn, J., Reinfelder, V., Stillfried, M., Heckmann, I., Scharf, A.K., Augeri, D.M., Cheyne, S.M., Hearn, A.J., Ross, J., Macdonald, D.W., Mathai, J., Eaton, J., Marshall, A.J., Semiadi, G., Rustam, R., Bernard, H., Alfred, R., Samejima, H., Duckworth, J.W., Breitenmoser-Wuersten, C., Belant, J.L., Hofer, H., Wilting, A. (2013): The importance of correcting for sampling bias in MaxEnt species distribution models. Divers. Distrib. 19: 1366-1379.
Lanfear, R., Calcott, B., Ho, S.Y.W., Guindon, S. (2012): PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29: 1695-1701.
Lanfear, R., Frandsen, P.B., Wright, A.M., Senfeld, T., Calcott, B. (2017): PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34: 772-773.
Li, J. (2023): iNaturalist observation 186300362. iNaturalist.org. Available online at: https://www.inaturalist.org/observations/186300362.
Li, J., Che, J., Bain, R.H., Zhao, E., Zhang, Y. (2008): Molecular phylogeny of Rhacophoridae (Anura): A framework of taxonomic reassignment of species within the genera Aquixalus, Chiromantis, Rhacophorus, and Philautus. Mol. Phylogenet. Evol. 48: 302-312.
Li, J.-T., Li, Y., Klaus, S., Rao, D.-Q., Hillis, D.M., Zhang, Y.-P. (2013): Diversification of rhacophorid frogs provides evidence for accelerated faunal exchange between India and Eurasia during the Oligocene. Proc. Natl. Acad. Sci. 110: 3441-3446.
Li, S., Dai, W., Wang, Z., Wu, Z., Wang, J. (2024): Detecting range shrinking from historical amphibian species occurrences under influence of human impacts: A case study using the Chinese giant salamander, Andrias davidianus. Ecol. Evol. 14: e70595.
Lin, Y.-S., Kam, Y.-C. (2008): Nest choice and breeding phenology of an arboreal-breeding frog, Kurixalus eifingeri (Rhacophoridae), in a bamboo forest. Zool. Stud. 47: 129-137.
Liu, C., Newell, G., White, M., Machunter, J. (2025): Improving the estimation of the Boyce index using statistical smoothing methods for evaluating species distribution models with presence‐only data. Ecography 2025: e07218.
Luedtke, J.A., Chanson, J., Neam, K., Hobin, L., Maciel, A.O., Catenazzi, A., Borzée, A., Hamidy, A., Aowphol, A., Jean, A., Sosa-Bartuano, Á., Fong G., A., De Silva, A., Fouquet, A., Angulo, A., Kidov, A.A., Muñoz Saravia, A., Diesmos, A.C., Tominaga, A., Shrestha, B., Gratwicke, B., Tjaturadi, B., Martínez Rivera, C.C., Vásquez Almazán, C.R., Señaris, C., Chandramouli, S.R., Strüssmann, C., Cortez Fernández, C.F., Azat, C., Hoskin, C.J., Hilton-Taylor, C., Whyte, D.L., Gower, D.J., Olson, D.H., Cisneros-Heredia, D.F., Santana, D.J., Nagombi, E., Najafi-Majd, E., Quah, E.S.H., Bolaños, F., Xie, F., Brusquetti, F., Álvarez, F.S., Andreone, F., Glaw, F., Castañeda, F.E., Kraus, F., Parra-Olea, G., Chaves, G., Medina-Rangel, G.F., González-Durán, G., Ortega-Andrade, H.M., Machado, I.F., Das, I., Dias, I.R., Urbina-Cardona, J.N., Crnobrnja-Isailović, J., Yang, J.-H., Jianping, J., Wangyal, J.T., Rowley, J.J.L., Measey, J., Vasudevan, K., Chan, K.O., Gururaja, K.V., Ovaska, K., Warr, L.C., Canseco-Márquez, L., Toledo, L.F., Díaz, L.M., Khan, M.M.H., Meegaskumbura, M., Acevedo, M.E., Napoli, M.F., Ponce, M.A., Vaira, M., Lampo, M., Yánez-Muñoz, M.H., Scherz, M.D., Rödel, M.-O., Matsui, M., Fildor, M., Kusrini, M.D., Ahmed, M.F., Rais, M., Kouamé, N.G., García, N., Gonwouo, N.L., Burrowes, P.A., Imbun, P.Y., Wagner, P., Kok, P.J.R., Joglar, R.L., Auguste, R.J., Brandão, R.A., Ibáñez, R., Von May, R., Hedges, S.B., Biju, S.D., Ganesh, S.R., Wren, S., Das, S., Flechas, S.V., Ashpole, S.L., Robleto-Hernández, S.J., Loader, S.P., Incháustegui, S.J., Garg, S., Phimmachak, S., Richards, S.J., Slimani, T., Osborne-Naikatini, T., Abreu-Jardim, T.P.F., Condez, T.H., De Carvalho, T.R., Cutajar, T.P., Pierson, T.W., Nguyen, T.Q., Kaya, U., Yuan, Z., Long, B., Langhammer, P., Stuart, S.N. (2023): Ongoing declines for the world’s amphibians in the face of emerging threats. Nature 622: 308-314.
Luo, T., Zhao, X., Lan, C., Li, W., Deng, H., Xiao, N., Zhou, J. (2023): Integrated phylogenetic analyses reveal the evolutionary, biogeographic, and diversification history of Asian warty treefrog genus Theloderma (Anura, Rhacophoridae). Ecol. Evol. 13: e10829.
Lv, Y.-Y., He, K., Klaus, S., Brown, R.M., Li, J.-T. (2018): A comprehensive phylogeny of the genus Kurixalus (Rhacophoridae, Anura) sheds light on the geographical range evolution of frilled swamp treefrogs. Mol. Phylogenet. Evol. 121: 224-232.
Lyu, Z., Li, M., Wang, G., Liu, G., Liu, M., Jiang, K., Jiang, D., Li, J. (2024): Taxonomic status of three recently proposed species of the genus Kurixalus (Anura, Rhacophoridae), with discussion on the ZooBank registrations for electronical publication. Asian Herpetol. Res. 15: 31-40.
Madhushanka, S., Manamendra-Arachchi, K. (2021): Morphometry and habitat selection of the mountain hourglass treefrog [Taruga eques (Günther, 1858)] (Amphibia, Rhacophoridae) in the Horton Plains National Park, Sri Lanka. Univ. Colombo Rev. 3: 64.
Messenger, K.R., Othman, S.N., Chuang, M.-F., Yang, Y., Borzée, A. (2022): Description of a new Kurixalus species (Rhacophoridae, Anura) and a northwards range extension of the genus. ZooKeys 1108: 15-49.
Mo, Q., Sun, T., Chen, H., Yu, G., Du, L. (2023): Biogeographic origin of Kurixalus (Anura, Rhacophoridae) on the east Asian islands and tempo of diversification within Kurixalus. Animals 13: 2754.
Nguyen, T.T., Matsui, M., Duc, H.M. (2014): A new tree frog of the genus Kurixalus (Anura: Rhacophoridae) from Vietnam. Curr. Herpetol. 33: 101-111.
Nguyen, T.V., Duong, T.V., Luu, K.T., Poyarkov, N.A. (2020): A new species of Kurixalus (Anura: Rhacophoridae) from northern Vietnam with comments on the biogeography of the genus. J. Nat. Hist. 54: 195-223.
Pan, T., Wang, H., Duan, S., Ali, I., Yan, P., Cai, R., Wang, M., Zhang, J., Zhang, H., Zhang, B., Wu, X. (2019): Historical population decline and habitat loss in a critically endangered species, the Chinese alligator (Alligator sinensis). Glob. Ecol. Conserv. 20: e00692.
Pang, C., Tang, S., Yu, G., Zhou, J.-J. (2024): First description of the female and morphological variations with range extension of Kurixalus lenquanensis (Anura, Rhacophoridae). Biodivers. Data J. 12: e130411.
Phillips, S.J., Anderson, R.P., Dudík, M., Schapire, R.E., Blair, M.E. (2017): Opening the black box: an open-source release of Maxent. Ecography 40: 887-893.
Proosdij, A.S.J. van, Sosef, M.S.M., Wieringa, J.J., Raes, N. (2016): Minimum required number of specimen records to develop accurate species distribution models. Ecography 39: 542-552.
QGIS.org (2023): QGIS. QGIS Association.
R Core Team (2022): R: A Language and environment for statistical computing. Vienna, Austria, Foundation for Statistical Computing.
Rhoden, C.M., Peterman, W.E., Taylor, C.A. (2017): Maxent-directed field surveys identify new populations of narrowly endemic habitat specialists. PeerJ 5: e3632.
Ronquist, F., Teslenko, M., Van Der Mark, P., Ayres, D.L., Darling, A., Höhna, S., Larget, B., Liu, L., Suchard, M.A., Huelsenbeck, J.P. (2012): MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61: 539-542.
Sarker, G.C., Wostl, E., Thammachoti, P., Sidik, I., Hamidy, A., Kurniawan, N., Smith, E.N. (2019): New species, diversity, systematics, and conservation assessment of the puppet toads of Sumatra (Anura: Bufonidae: Sigalegalephrynus). Zootaxa 4679: 365-391.
Shin, Y., Min, M., Borzée, A. (2021): Driven to the edge: Species distribution modeling of a clawed salamander (Hynobiidae: Onychodactylus koreanus) predicts range shifts and drastic decrease of suitable habitats in response to climate change. Ecol. Evol. 11: 14669-14688.
Trifinopoulos, J., Nguyen, L.-T., von Haeseler, A., Minh, B.Q. (2016): W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44: W232–W235.
Vignali, S., Barras, A.G., Arlettaz, R., Braunisch, V. (2020): SDMtune: An R package to tune and evaluate species distribution models. Ecol. Evol. 10: 11488-11506.
Warren, D.L., Seifert, S.N. (2011): Ecological niche modeling in Maxent: the importance of model complexity and the performance of model selection criteria. Ecol. Appl. 21: 335-342.
Wilkinson, J.A., Drewes, R.C., Tatum, O.L. (2002): A molecular phylogenetic analysis of the family Rhacophoridae with an emphasis on the Asian and African genera. Mol. Phylogenet. Evol. 24: 265-273.
Wu, S.-P., Huang, C.-C., Tsai, C.-L., Lin, T.-E., Jhang, J.-J., Wu, S.-H. (2016): Systematic revision of the Taiwanese genus Kurixalus members with a description of two new endemic species (Anura, Rhacophoridae). ZooKeys 557: 121-153.
Xu, W., Wu, Y.-H., Zhou, W.-W., Chen, H.-M., Zhang, B.-L., Chen, J.-M., Xu, W., Rao, D.-Q., Zhao, H., Yan, F., Yuan, Z., Jiang, K., Jin, J.-Q., Hou, M., Zou, D., Wang, L.-J., Zheng, Y., Li, J.-T., Jiang, J., Zeng, X.-M., Chen, Y., Liao, Z.-Y., Li, C., Li, X.-Y., Gao, W., Wang, K., Zhang, D.-R., Lu, C., Yin, T., Ding, Z., Zhao, G.-G., Chai, J., Zhao, W.-G., Zhang, Y.-P., Wiens, J.J., Che, J. (2024): Hidden hotspots of amphibian biodiversity in China. Proc. Natl. Acad. Sci. 121: e2320674121.
Yu, G., Hui, H., Rao, D., Yang, J. (2018): A new species of Kurixalus from western Yunnan, China (Anura, Rhacophoridae). ZooKeys 770: 211-226.
Yu, G., Rao, D., Matsui, M., Yang, J. (2017a): Coalescent-based delimitation outperforms distance-based methods for delineating less divergent species: the case of Kurixalus odontotarsus species group. Sci. Rep. 7: 16124.
Yu, G., Wang, J., Hou, M., Rao, D., Yang, J. (2017b): A new species of the genus Kurixalus from Yunnan, China (Anura, Rhacophoridae). ZooKeys 694: 71-93.
Yu, G.-H., Du, L.-N., Wang, J.-S., Rao, D.-Q., Wu, Z.-J., Yang, J.-X. (2020): From mainland to islands: colonization history in the tree frog Kurixalus (Anura: Rhacophoridae). Curr. Zool. 66: 667-675.
Zanaga, D., Van De Kerchove, R., De Keersmaecker, W., Souverijns, N., Brockmann, C., Quast, R., Wevers, J., Grosu, A., Paccini, A., Vergnaud, S., Cartus, O., Santoro, M., Fritz, S., Georgieva, I., Lesiv, M., Carter, S., Herold, M., Li, L., Tsendbazar, N.-E., Ramoino, F., Arino, O. (2021): ESA WorldCover 10 m 2020 v100. Zenodo. https://doi.org/10.5281/ZENODO.5571936.
Zeng, J., Wang, J.-S., Yu, G.-H., Du, L.-N. (2021): A new species of Kurixalus (Anura, Rhacophoridae) from Guizhou, China. Zool. Res. 42: 227-233.
Zhao, E., Wang, L., Shi, H., Wu, G., Zhao, H. (2005): Chinese rhacophorid frogs and description of a new species of Rhacophorus. Sichuan J. Zool. 24: 297-300.
Zhu, B., Wang, J., Sun, Z., Yang, Y., Wang, T., Brauth, S.E., Tang, Y., Cui, J. (2017): Competitive pressures affect sexual signal complexity in Kurixalus odontotarsus: insights into the evolution of compound calls. Biol. Open 6: 1913-1918.
Zhu, G., Qiao, H. (2016): Effect of the Maxent model’s complexity on the prediction of species potential distributions. Biodivers. Sci. 24: 1189-1196.